Modules of co-regulated genes have been identified throughout the ImmGen data, with prediction of transcriptional regulators via the Ontogenet algorithm (Jojic et al, 2013). Many aspects of these metadata can be explored here.
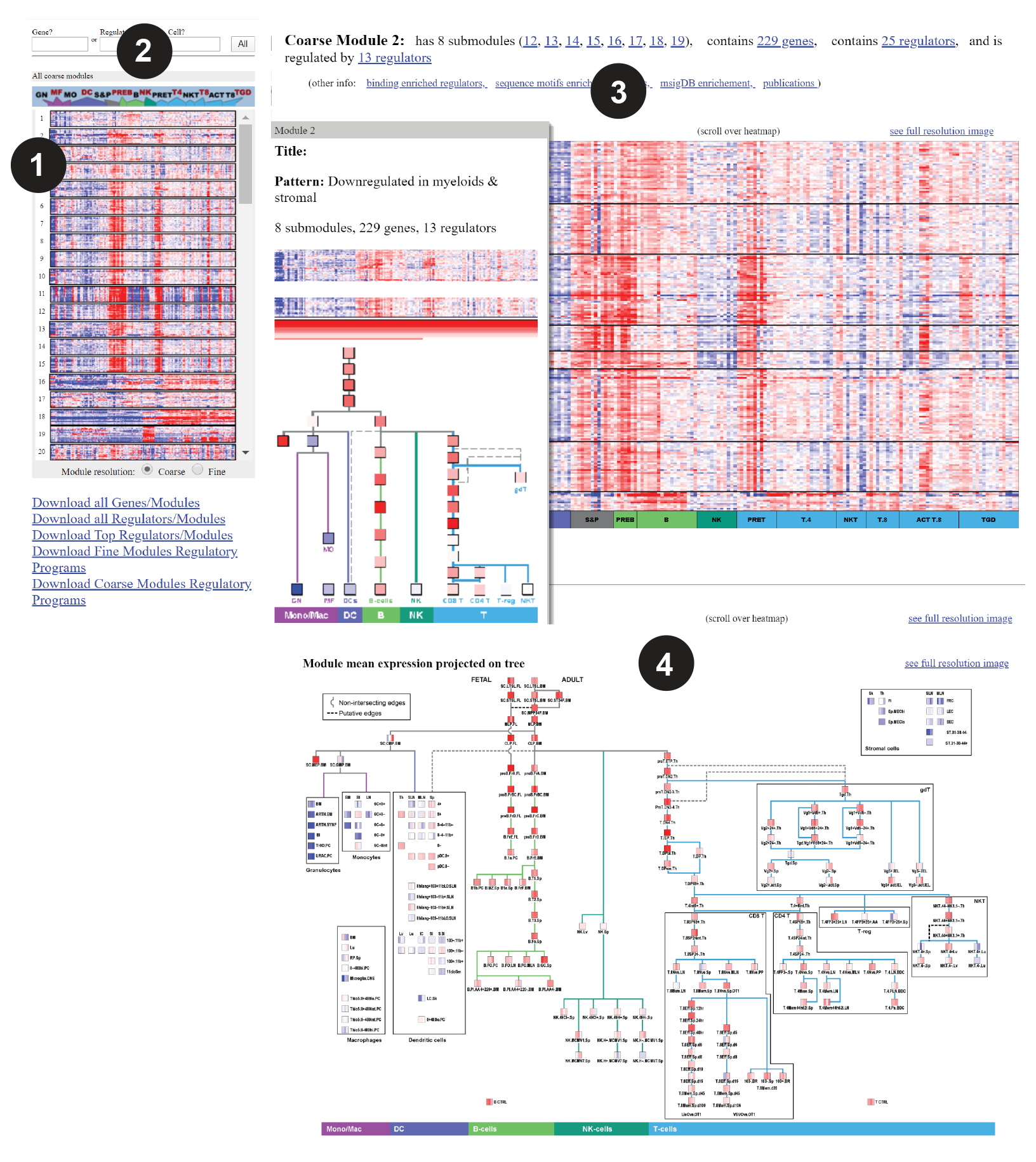
The “Exploration bar” at left shows thumbnail representations of the modules’ expression pattern across ImmGen cell-types (1), in coarse or fine modules.
The search boxes (2) bring up modules that contain a specific geneor are controlled by a predicted activator/repressor. Click on a thumbnail to retrieve full information on the module: interactive heat maps of the module’s and its regulators’ expression (mouse-over for details), the matrix of regulator weights, and the module’s mean expression projected on the ImmGen differentiation tree. Tables of module membership, of activators and repressors are accessed at top (3).
Additional information related to the module’s genes enrichment in sequence motifs, in msigDB genesets, or to publications associating at least two of the module members can be accessed (4).

