 |
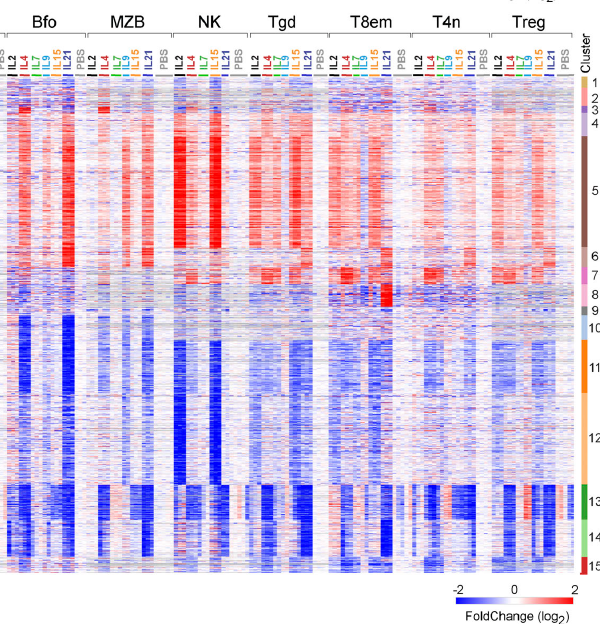
The interweaved signatures of common-gamma-chain cytokines across immunologic lineages.
Baysoy A, Seddu K, Salloum T, Dawson CA, Lee JJ, Yang L, Gal-Oz S, Ner-Gaon H, Tellier J, Millan A, Sasse A, Brown B, Lanier LL, Shay T, Nutt S, Dwyer D, Benoist C; Immunological Genome Project Consortium.
J Exp Med. 2023 Jul 3;220(7):e20222052.
Baysoy A, Seddu K, Salloum T, Dawson CA, Lee JJ, Yang L, Gal-Oz S, Ner-Gaon H, Tellier J, Millan A, Sasse A, Brown B, Lanier LL, Shay T, Nutt S, Dwyer D, Benoist C; Immunological Genome Project Consortium.
J Exp Med. 2023 Jul 3;220(7):e20222052.
| Pubmed |
|
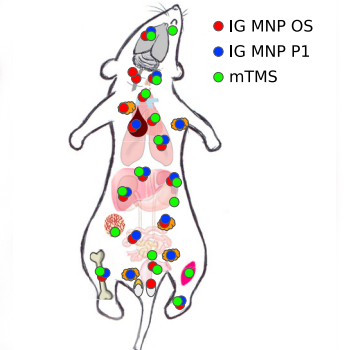 |
Network analysis of large-scale ImmGen and Tabula Muris datasets highlights metabolic diversity of tissue mononuclear phagocytes.
Gainullina A, Mogilenko D, Huang LH,Todorov H, Narang V, Kim KW, Yng LS,
Kent A, Baosen Jia B, Seddu K,Krchma K,Wu J, Crozat K, Tomasello E,
Dress R, See P, Scott C, Gibbings S, Bajpai G,
Desai JV,Maier B,This S,Wang P, Vargas Aguilar S,
Poupel L,Dussaud S, Zhou TA,Angeli V, Blander JM, Choi K, Dalod M, Dzhagalov I, Gautier EL,
Jakubzick C, Lavine K, Lionakis MS, Paidassi H, Michael H, Sieweke MH,
Ginhoux F, Guilliams M, Benoist C, Merad M, Randolph G, Sergushichev A, Artyomov M; ImmGen Consortium. Cell Rep. 2023 Jan 27;42(2):112046.
|
Pubmed |
|
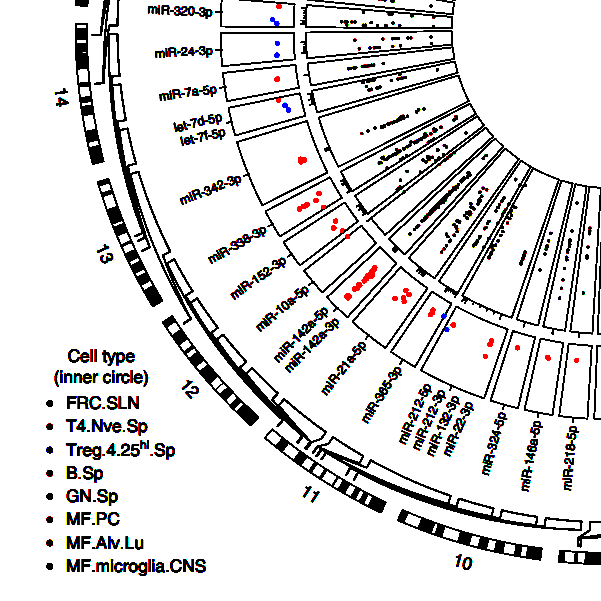 |
A microRNA expression and regulatory element activity atlas of the mouse immune system.
Rose SA, Wroblewska A, Dhainaut M, Yoshida H, Shaffer JM, Bektesevic A, Ben-Zvi B,
Rhoads A, Kim EY, B, Lavin Y, Merad M, Buenrostro JD, Brown BD, Immunological Genome Consortium. Nat Immunol. 2021 Jul;22(7):914-927.
|
Pubmed |
|
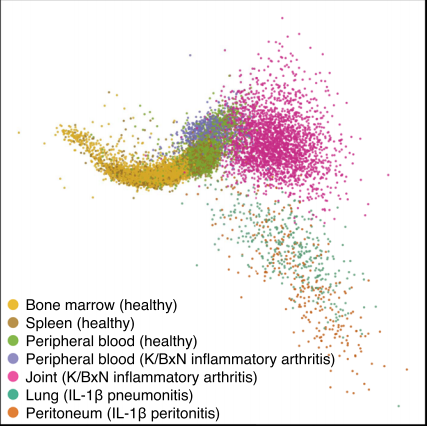 |
The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments
Grieshaber-Bouyer R, Radtke FA, Cunin P, Stifano G, Levescot A, Vijaykumar B, Nelson-Maney N, Blaustein RB, Monach P, Nigrovic PA, ImmGen Consortium
Nat Commun. 2021 May 17;12(1):2856.
|
Pubmed |
|
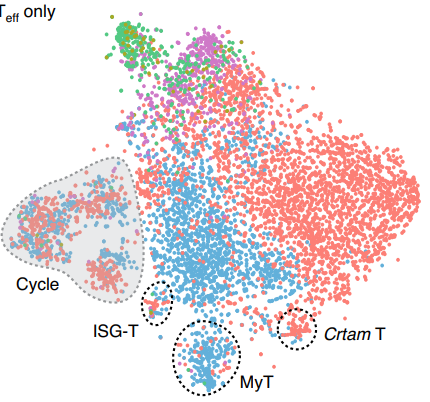 |
Gut CD4+ T cell phenotypes are a continuum molded by microbes, not by TH archetypes.
Kiner E, Willie E, Vijaykumar B, Chowdhary K, Schmutz H, Chandler J, Schnell A, Thakore PI, LeGros G, Mostafavi S, Mathis D, Benoist C; Immunological Genome Project Consortium.
Nat Immunol. 2021 Feb;22(2):216-228.
|
Pubmed |
|
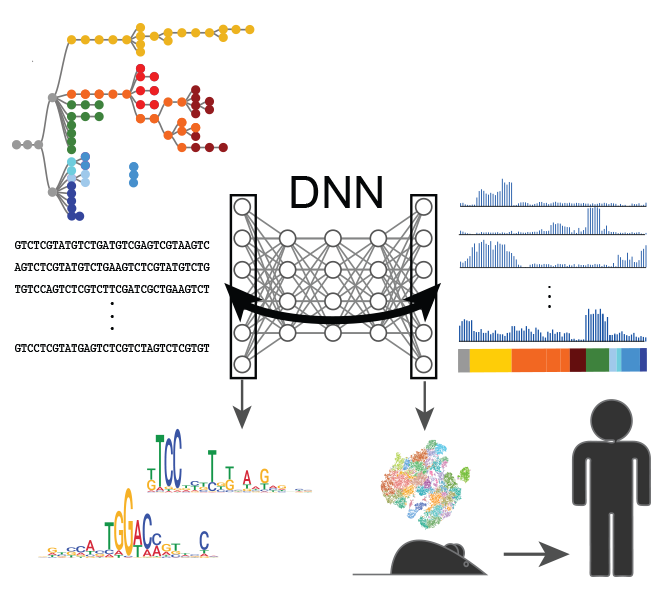 |
Deep learning of immune cell differentiation.
Maslova A, Ramirez RN, Ma K, Schmutz H, Wang C, Fox C, Ng B, Benoist C, Mostafavi S; Immunological Genome Project.
Proc Natl Acad Sci U S A. 2020 Oct 13;117(41):25655-25666.
|
Pubmed |
|
 |
ImmGen at 15
Immunological Genome Project.
Nat Immunol. 2020 Jul;21(7):700-703.
|
Pubmed |
|
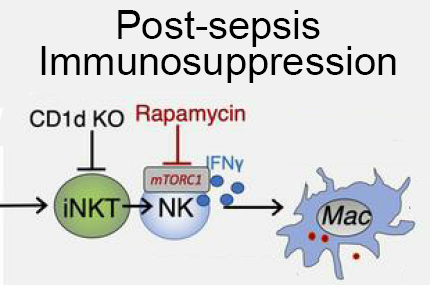 |
Post-sepsis immunosuppression depends on NKT cell regulation of mTOR/IFN-γ in NK cells.
Kim E,Ner-Gaon H, Varon J, Cullen AM, Guo J, Choi J, Barragan-Bradford D, Higuera A, Pinilla-Vera M, Short SA,
Arciniegas-Rubio A
, Tamura T, Leaf DE, Baron RM, Shay T
, Brenner MB.
J Clin Invest. 2020 Jun 1;130(6):3238-3252. |
Pubmed |
|
 |
ImmGen report: sexual dimorphism in the immune system transcriptome.
Gal-Oz ST, Maier B, Yoshida H, Seddu K, Elbaz N, Czysz C, Zuk O, Stranger BE, Ner-Gaon H, Shay T. Nat Commun. 2019 Sep 20;10(1):4295.
|
Pubmed |
|
 |
The cis-Regulatory Atlas of the Mouse Immune System.
Yoshida H, Lareau C, Ramirez R, Rose S, Maier B, Wroblewska A, Desland F, Chudnovskiy A, Mortha A, Dominguez C, Tellier J, Kim E, Dwyer D, Shinton S, Nabekura T, Qi Y, Yu B, Robinette M, Kim KW, Wagers A, Rhoads A, Nutt S, Brown B, Mostafavi S, Buenrostro J, Benoist, C, The Immunological Genome Project. 2019, Cell 176, 897–912.
|
Pubmed |
|
 |
Expression profiling of constitutive mast cells reveals a unique identity within the immune system.
Dwyer DF, Barrett NA, Austen KF Immunological Genome Project Consortium. Nat Immunol. 2016 Jul;17(7):878-87.
|
Pubmed |
|
 |
Parsing the Interferon Transcriptional Network and Its Disease Associations.
Mostafavi S, Yoshida H, Moodley D, LeBoité H, Rothamel K, Raj T, Ye CJ, Chevrier N, Zhang SY, Feng T, Lee M, Casanova JL, Clark JD, Hegen M, Telliez JB, Hacohen N, De Jager PL, Regev A, Mathis D, Benoist C; Immunological Genome Project Consortium. Cell. 2016 Jan 28;164(3):564-78.
|
Pubmed |
|
 |
Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets.
Robinette ML, Fuchs A, Cortez VS, Lee JS, Wang Y, Durum SK, Gilfillan S, Colonna M; the Immunological Genome Consortium. Nat Immunol. 2015 Mar;16(3):306-17.
|
Pubmed |
|
 |
Gene expression during the generation and activation of mouse neutrophils: implication of novel functional and regulatory pathways.
Ericson JA, Duffau P, Yasuda K, Ortiz-Lopez A, Rothamel K, Rifkin IR, Monach PA, ImmGen Consortium. PLoS One. 2014 Oct 3;9(10):e108553.
|
Pubmed |
|
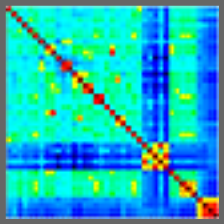 |
Variation and Genetic Control of Gene Expression in Primary Immunocytes across Inbred Mouse Strains.
Mostafavi S, Ortiz-Lopez A, Bogue MA, Hattori K, Pop C, Koller D, Mathis D, Benoist C; The Immunological Genome Consortium, Blair DA, Dustin ML, Shinton SA, Hardy RR, Shay T, Regev A, Cohen N, Brennan P, Brenner M, Kim F, Rao TN, Wagers A, Heng T, Ericson J, Rothamel K, Ortiz-Lopez A, Mathis D, Benoist C, Kreslavsky T, Fletcher A, Elpek K, Bellemare-Pelletier A, Malhotra D, Turley S, Miller J, Brown B, Merad M, Gautier EL, Jakubzick C, Randolph GJ, Monach P, Best AJ, Knell J, Goldrath A, Jojic V, Koller D, Laidlaw D, Collins J, Gazit R, Rossi DJ, Malhotra N, Sylvia K, Kang J5, Bezman NA, Sun JC, Min-Oo G, Kim CC, Lanier LL. J Immunol. 2014 Nov 1;193(9):4485-96.
|
Pubmed |
|
 |
Immunological Genome Project and systems immunology.
Shay T, Kang J. Trends Immunol. 2013 Dec;34(12):602-9.
|
Pubmed |
|
 |
Beyond the transcriptome: completion of act one of the Immunological Genome Project.
Kim CC, Lanier LL. Curr Opin Immunol. 2013 Oct;25(5):593-7.
|
Pubmed |
|
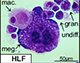 |
Transcriptome analysis identifies regulators of hematopoietic stem and progenitor cells.
Gazit R, Garrison BS, Rao TN, Shay T, Costello J, Ericson J, Kim F, Collins JJ, Regev A, Wagers AJ, Rossi DJ; Immunological Genome Project Consortium. Stem Cell Reports. 2013 Aug 15;1(3):266-80.
|
Pubmed |
|
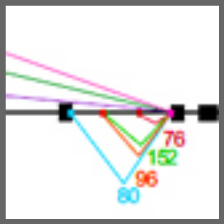 |
Differential splicing across immune system lineages.
Ergun A, Doran G, Costello JC, Paik HH, Collins JJ, Mathis D, Benoist C; ImmGen Consortium. Proc Natl Acad Sci U S A. 2013 Aug 27;110(35):14324-9.
|
Pubmed |
|
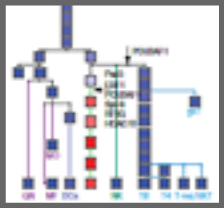 |
Identification of transcriptional regulators in the mouse immune system.
Jojic V, Shay T, Sylvia K, Zuk O, Sun X, Kang J, Regev A, Koller D; the Immunological Genome Project Consortium. Nat Immunol. 2013 Jun;14(6):633-43.
|
Pubmed |
|
 |
The transcriptional landscape of αβ T cell differentiation.
Mingueneau M, Kreslavsky T, Gray D, Heng T, Cruse R, Ericson J, Bendall S, Spitzer M, Nolan G, Kobayashi K, von Boehmer H, Mathis D, Benoist C; the Immunological Genome Consortium. Nat Immunol. 2013 Jun;14(6):619-32.
|
Pubmed |
|
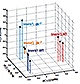 |
A network of high-mobility group box transcription factors programs innate interleukin-17 production.
Malhotra N, Narayan K, Cho OH, Sylvia KE, Yin C, Melichar H, Rashighi M, Lefebvre V, Harris JE, Berg LJ, Kang J; the Immunological Genome Project Consortium. Immunity. 2013 Apr 18;38(4):681-93.
|
Pubmed |
|
 |
Transcriptional insights into the CD8+ T cell response to infection and memory T cell formation.
Best JA, Blair DA, Knell J, Yang E, Mayya V, Doedens A, Dustin ML, Goldrath AW; The Immunological Genome Project Consortium, Nat Immunol. 2013 Apr;14(4):404-12.
|
Pubmed |
|
 |
Conservation and divergence in the transcriptional programs of the human and mouse immune systems.
Shay T, Jojic V, Zuk O, Rothamel K, Puyraimond-Zemmour D, Feng T, Wakamatsu E, Benoist C, Koller D, Regev A; the Immunological Genome Consortium. Proc Natl Acad Sci U S A. 2013 Feb 19;110(8):2946-51.
|
Pubmed |
|
 |
Decoding dendritic cell function through module and network analysis.
Pandey G, Cohain A, Miller J, Merad M. J Immunol Methods. 2013 Jan 31;387(1-2):71-80.
|
Pubmed |
|
 |
Shared and distinct transcriptional programs underlie the hybrid nature of iNKT cells.
Cohen NR, Brennan PJ, Shay T, Watts GF, Brigl M, Kang J, Brenner MB; the Immunological Genome Consortium. Nat Immunol. 2013 Jan;14(1):90-9.
|
Pubmed |
|
 |
Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages.
Gautier EL, Shay T, Miller J, Greter M, Jakubzick C, Ivanov S, Helft J, Chow A, Elpek KG, Gordonov S, Mazloom AR, Ma'ayan A, Chua WJ, Hansen TH, Turley SJ, Merad M, Randolph GJ; the Immunological Genome Consortium. Nat Immunol. 2012 Sep 30;13(11):1118-1128.
|
Pubmed |
|
 |
Molecular definition of the identity and activation of natural killer cells.
Bezman NA, Kim CC, Sun JC, Min-Oo G, Hendricks DW, Kamimura Y, Best JA, Goldrath AW, Lanier LL; the Immunological Genome Consortium. Nat Immunol. 2012 Aug 19;13(10):1000-1009.
|
Pubmed |
|
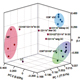 |
Deciphering the transcriptional network of the dendritic cell lineage.
Miller JC, Brown BD, Shay T, Gautier EL, Jojic V, Cohain A, Pandey G, Leboeuf M, Elpek KG, Helft J, Hashimoto D, Chow A, Price J, Greter M, Bogunovic M, Bellemare-Pelletier A, Frenette PS, Randolph GJ, Turley SJ, Merad M; the Immunological Genome Consortium. Nat Immunol. 2012 Jul 15;13(9):888-899.
|
Pubmed |
|
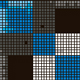 |
Transcriptional profiling of stroma from inflamed and resting lymph nodes defines immunological hallmarks.
Malhotra D, Fletcher AL, Astarita J, Lukacs-Kornek V, Tayalia P, Gonzalez SF, Elpek KG, Chang SK, Knoblich K, Hemler ME, Brenner MB, Carroll MC, Mooney DJ, Turley SJ & the Immunological Genome Project Consortium. Nat Immunol. 2012 Apr 1;13(5):511-518.
|
Pubmed |
|
 |
Intrathymic programming of effector fates in three molecularly distinct γδ T cell subtypes.
Narayan K, Sylvia KE, Malhotra N, Yin CC, Martens G, Vallerskog T, Kornfeld H, Xiong N, Cohen NR, Brenner MB, Berg LJ, Kang J; The Immunological Genome Project Consortium. Nat Immunol. 2012 Apr 1;13(5):499-510.
|
Pubmed |
|
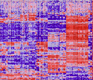 |
Transcriptomes of the B and T lineages compared by multiplatform microarray profiling.
Painter MW, Davis S, Hardy RR, Mathis D, Benoist C; Immunological Genome Project Consortium. J Immunol. 2011 Mar 1;186(5):3047-57.
|
Pubmed |
|
 |
The Immunological Genome Project: networks of gene expression in immune cells.
Heng TS, Painter MW; Immunological Genome Project Consortium. Nat Immunol. 2008 Oct;9(10):1091-4.
|
Pubmed |
|















